The FeMo-cofactor and classical and quantum computing
Recently, my coworkers and I put out a preprint “Classical solution
of the FeMo-cofactor model to chemical accuracy and its
implications’’ (Zhai et al. 2026). It is a bit
unusual to write commentary on one’s own scientific article. However, in
this case, given the many inquiries I have had about the work in the
context of quantum computing, many of which have contained similar
questions (and often similar misunderstandings), I thought it would be
useful to provide some perspective that we could not provide in the
original preprint, in an informal manner.
I will start with some background on the FeMo-cofactor (FeMo-co).
This cofactor is the reaction center of nitrogenase, an enzyme found in
certain soil-dwelling bacteria. Nitrogenase’s claim to fame is that it
converts atmospheric dinitrogen, which is held together by a strong N-N
triple bond, into a reduced form (ammonia) which can then be taken up by
plants and thereby be passed onto the rest of the living biomass. In
terms of incorporating nitrogen into biomass, nitrogenase is believed to
be responsible for about 2/3 of
biological nitrogen, with the remainder coming from fertilizers. Because
it plays this critical role, it is sometimes referred to as the enzyme
that feeds the planet.
The chemistry of how dinitrogen is reduced at the FeMo-cofactor is still largely unknown. The basic stoichiometry of the reaction is often written as
but this just a sketch of the process. In particular, the above equation contains, nominally, a large number of molecular reactants, and clearly they do not all just come together in a bang! The role of the cofactor, and the enzyme more generally, is to coordinate the protons, electrons, biological energy source (ATP), and the dinitrogen molecule, into a sequence of well-defined steps, known as the reaction mechanism. Since the work of Lowe and Thorneley (Thorneley and Lowe 1984), the most common proposal for the nitrogenase reaction mechanism contains 8 intermediate steps (corresponding roughly to 8 sequential proton and electron additions). However, due to the difficulty in isolating the intermediate states of FeMo-co, as well as challenges in using experimental probes to deduce what these states are, the Lowe-Thorneley cycle still remains an unproven hypothesis. Biochemists, spectroscopists, as well as a few theoretical quantum chemists, are today actively engaged in observing, computing, deducing (and arguing about) the nitrogenase mechanism (Jiang and Ryde 2023; Lancaster et al. 2011; Einsle and Rees 2020; Badding et al. 2023; Thorhallsson et al. 2019).
So how did nitrogenase become so widely discussed in the setting of
quantum computing? In 2016, an article “Elucidating reaction mechanisms
on quantum computers’’, that has since become one of the most cited
papers in the nitrogenase field, arguably started this all (Reiher et al.
2017). The article included a number of proposals, including (1)
that the ‘promise of exponential speedups for the electronic structure
problem’ could be applied to elucidate the nitrogenase reaction
mechanism that had so far proved intractable for classical computation,
and (2) that solving this problem would be an example of how quantum
simulation could be ‘scientifically and economically impactful’.
(Similar proposals can also be found repeated in less technical language
and settings, see e.g. ‘Why do
we want a quantum computer’). An important technical contribution of
the article was to provide a detailed quantum resource estimate for a
simulation of chemistry. The problem statement was to compute the
ground-state energy of a specific ‘54 orbital’ (108 qubit) model of
FeMo-co, to an accuracy of 1 kcal/mol, referred to as chemical accuracy.
It is important to note the word ‘model’ in the problem statement.
Electrons move in continuous space, and thus quantum chemical
Hamiltonians are formulated in the continuum, while quantum computation
requires discretization of this space. This discretization, in terms of
a so-called active space set of orbitals that the electrons can
variously occupy, constitutes the model. We will return to the
definition of the model below. By compiling a Trotter-Suzuki
implementation of the quantum phase estimation algorithm within a
fault-tolerant resource model for their specific FeMo-co model
Hamiltonian, Ref. (Reiher et al. 2017) provided
a T-gate resource estimate. Combined with some assumptions about the
quantum architecture, this provided perhaps the first concrete time-cost
to solve an interesting chemistry problem on a quantum computer. This
work has since served as an inspiration for many subsequent quantitative
resource estimation efforts in the quantum computing for chemistry
field.
Before proceeding further in this story, it is worth examining the
two key propositions made in Ref. (Reiher et al. 2017). I start
with the question of exponential speedup. Quantum algorithms for the
ground-state energy, such as quantum phase estimation, essentially
perform a projective measurement of the energy (encoded in a phase).
Thus, it is essential to prepare a good initial state, i.e. with large
overlap with the desired eigenstate, to measure the correct energy.
This, however, is a strong constraint, if we are seeking asymptotically
large quantum advantage. For example, if such an initial state is first
determined classically, as is often suggested, then exponential quantum
advantage in a given problem requires that finding good classical
guesses is easy, while improving them classically to fully solve the
problem becomes exponentially hard as the problem size increases.
Unfortunately, convincing evidence that chemically relevant electronic
structure problems, including the problem of cofactor electronic
structure exemplified by FeMo-co, fall into this category has not yet
been found, as discussed in detail in Refs. (Lee et al. 2023; Chan
2024).
The second proposition, that elucidating the reaction mechanism of
nitrogenase will lead to a transformative societal impact, is similarly
nuanced. The claim originates in the observation that the competing
industrial process for fertilizer production via nitrogen reduction,
namely, the Haber-Bosch process, takes place at high temperatures and
pressures and consumes a significant percentage of the world’s energy.
Bacteria, on the other hand, can do this process at room
temperature.
While it is true that the nitrogenase enzyme functions at ambient
temperature and pressure, it is simply false that it consumes much less
energy. This is because the large amount of energy required for nitrogen
fixation mainly originates from thermodynamics, i.e. one needs energy to
break the strong nitrogen triple bond. In fact, taking into account the
physiological conditions and the ATP cost, bacteria arguably expend
more energy to reduce ammonia (Chan 2024) than a modern efficient
industrial implementation of the Haber-Bosch process. Thus the real hope
behind trying to understand the nitrogenase mechanism in the context of
societal impact is that we may one day engineer a variant of it with
more desirable properties, e.g. with higher turnover, or with a lower
carbon footprint, or which is more selective for nitrogen reduction.
Whether this is actually possible remains to be seen, and certainly
requires much more than knowing the ground-state of FeMo-co, or even the
full reaction mechanism.
I now return to the question of FeMo-cofactor models. Ref. (Reiher et al.
2017) introduced a particular cofactor model, which I will refer
to it as RWST, following the names of the authors. As we soon found out,
simulating the ground-state of the RWST model was actually very easy
classically, and in fact (as reported in (Li et al. 2019)) could be done
using standard quantum chemistry methods with a few hours of calculation
on a laptop. This was because although the RWST model was a 108 qubit
model, and (in the worst case) a 108 qubit ground-state cannot be stored
classically, the RSWT model Hamiltonian was constructed in such a way to
not capture any of the difficult features of the FeMo-cofactor
ground-state. This highlights the importance of not assuming worst case
complexity about physical problems!
What makes the electronic structure of the FeMo-cofactor (relatively)
complicated is the presence of many ‘unpaired’ electrons. In simple
molecules, we can describe the ground-state as one where all the
electrons sit in pairs in orbitals. Since an orbital can only carry a
pair of electrons at a time, the ground-state is simply described by
filling the lowest energy orbitals with pairs. However, in molecules
with transition metals, there are typically ‘unpaired’ electrons
(so-called open-shells), and then we need to consider whether and how
they pair up, which orbitals are singly versus doubly occupied, and so
on. The RSWT model ground-state had no unpaired electrons! It was
therefore unrepresentatively easy to solve for the ground state
classically.
Because of the problems with the RWST model, my group formulated a
more suitable 76 orbital/152 qubit model of FeMo-co in Ref. (Li et al. 2019),
which I will refer to as the LLDUC model, again by the names of the
authors. Although the LLDUC model is still a significant truncation of
the true electronic structure of FeMo-co, we verified that it contains
the correct open-shell character of the cofactor, and thus has a
‘representative’ complexity in its ground-state. Since we published the
LLDUC model, it has become the most common benchmark model of FeMo-co
used in quantum resource estimates for new quantum chemistry
ground-state algorithms (Wan
et al. 2022; Berry et al. 2019; Luo and Cirac 2025; Low et al.
2025).
This brings me now to the recent work in Ref. (Zhai et al. 2026), where, through
a sequence of classical calculations, we could produce a classical
estimate of the ground-state energy of the LLDUC model to chemical
accuracy. How was this achieved?
Classical electronic structure methods (aside from exact
diagonalization) are heuristic algorithms. Much like quantum algorithms,
they implicitly or explicitly start from an initial state. In chemical
applications, this can be viewed as a product state or set of product
states: for tensor network algorithms, such as the density matrix
renormalization group (when not considering topological order) this is
the set of states (specified by
the underlying basis) which connect to the space of slightly entangled
states with that the
algorithm naturally explores. In coupled cluster methods, this is the
initial reference state to which excitations are applied. Although many
classical heuristics are exact with exponential effort, e.g. by
increasing the bond dimension in
a tensor network or excitation level in coupled cluster theory, in
practical computational chemistry, classical heuristics are used with
the assumption that so long as the initial state is chosen
appropriately, they will converge rapidly to the true ground-state
without exponential effort. I analyze this heuristic working assumption
in Ref. (Chan
2024) where I name it the classical heuristic cost conjecture.
However, finding the good classical initial state is an NP hard problem,
and this is often the crux of where the challenge in simulation actually
lies.
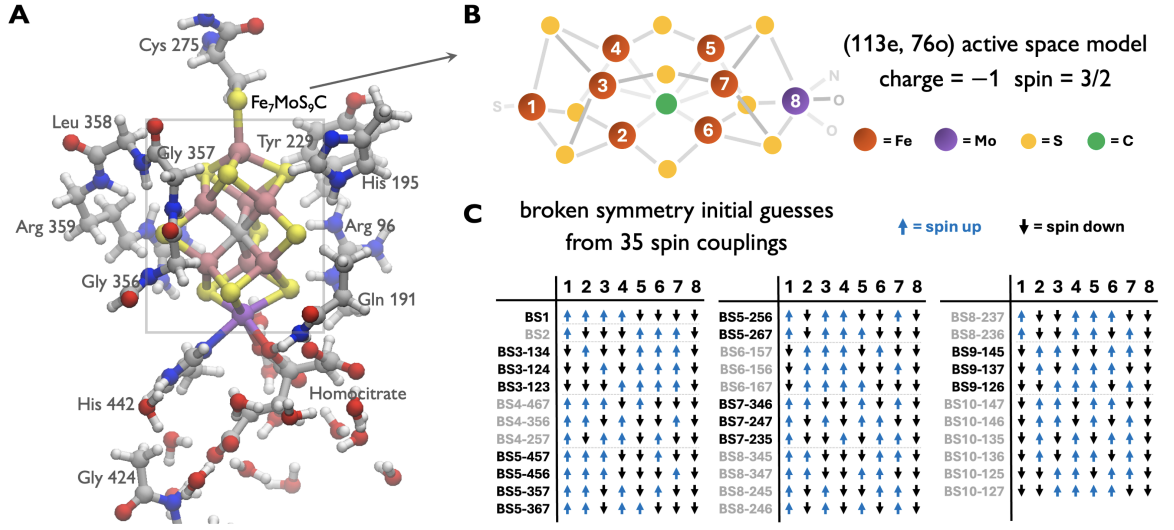
In FeMo-co, unlike in simpler molecules, it is not at all obvious
what product state to start from. To address this, in Ref. (Zhai et al.
2026), we devised an enumeration and filtering protocol. The
relevant manifold arises from the orbital and spin degrees of freedom of
the Fe ions: which Fe orbitals are occupied, by how many electrons, and
with which spins. One technical point is that the resulting product
states do not generally conserve the global spin symmetry. However, as recognized
by Anderson decades ago, for magnetic order in large systems, the
eigenstates can be chosen to break symmetry due to the tower of symmetry
preserving eigenstates at an energy scale of (where
is the system volume). For a finite
energy resolution we can equally use a broken symmetry description
of the states, an example of the fragility of entanglement effects in
physical systems.
Because applying the highest level of classical approximation to all
enumerated product states was far too expensive, we used a filtering
funnel, where product states were ranked at different levels of theory,
passing promising candidates to higher levels of classical computation.
In the end, the final most accurate calculations were performed on only
3 candidates, which we deduced to all be essentially degenerate to
within chemical accuracy.
There are other important technical details in Ref. (Zhai et al.
2026) which I have not mentioned: the use of unrestricted
orbitals, the systematic extrapolations to obtain the final energies and
estimated errors, and the benchmarking required to be confident about
the protocol. However, recognizing that the FeMo-co ground-state problem
could be reduced essentially to a ranking problem was the essence of
what made the estimate possible.
From a chemical and biochemical perspective, computing the
ground-state energy of a model to some specified accuracy – even
chemical accuracy – is a highly artificial target. Most chemical
calculations that have an impact on our understanding never achieve or
even target chemical accuracy in the total energy. In addition,
chemistry does not depend on the the total energy, but the relative
energy of different chemical configurations, which typically differ only
by changes in the bonding and
chemistry.
The main take-home from our work then is that there is nothing
especially mysterious about FeMo-co’s electronic structure. The story of
the FeMo-co ground-state is not one of multiconfigurational electronic
structure (i.e. where the states are not at all close to product
states), but one of multiple configurations (i.e. many competing product
states). Indeed, this is basically how nitrogenase chemists have long
reasoned about the electronic structure of iron-sulfur clusters and
FeMo-co (Lovell et al. 2001;
Yamaguchi et al. 1990). Our work thus now provides extensive and
rigorous numerical support for this picture.
Because of this simplicity, the full richness of classical quantum
chemistry methods can now be brought to bear on FeMo-co electronic
structure beyond the LLDUC model. Assuming the model already captures
the qualitative complexity of the cluster’s electronic structure, we
expect such investigations to provide quantitative corrections to the
picture we have obtained. We took some initial steps to confirm this in
our manuscript, considering larger orbital spaces, the effect of protein
fluctuations, and the interpretation of certain spectroscopies. In the
future, connecting these simulations to more spectroscopic measurements
will be an exciting possibility. In addition, now that the electronic
structure is on a conceptually sound footing, we have a foundation to
support the central question of resolving the reaction mechanism. This
opens up a whole new set of scientific challenges associated with
observing reactions on extremely slow timescales.
Because of the success of classical heuristic methods for this
problem, one may naturally wonder what these results mean for the
application of quantum computers in chemistry. Here I address some
commonly asked questions.
Is the classical simulation of the LLDUC model a ‘last hurrah’ for classical methods?
I have seen the analogy drawn between the FeMo-co result and the classical tensor network simulations for random circuit sampling experiments. In that case, while the famous Google Sycamore experiment (Arute et al. 2019) could be replicated by classical tensor network simulations (Gray and Kourtis 2021), subsequent improvements in quantum processors, soon led to random circuit sampling experiments outpacing the capabilities of classical simulations.
However, the situation here is quite different. There is strong
evidence that generating samples from a random quantum circuit (without
noise) is actually exponentially hard to do using a classical algorithm,
and indeed, the classical simulations used for the task were (mostly)
brute force simulations with exponential cost in circuit size. In
contrast, the theoretical support for exponential quantum advantage in
the FeMo-co problem is much weaker, and as an empirical fact, most of
the methods used in the FeMo-co simulation (namely the coupled cluster
methods for a given excitation level) are polynomial cost algorithms.
Since a similar simulation strategy has also been successfully applied
across the series of 2, 4, and 8 metal iron-sulfur complexes (Sharma
et al. 2014; Li et al. 2019; Zhai et al. 2023, 2026), we have no
reason to expect a radically different situation if we consider larger
analogous complexes in this series.
And in any case, chemistry does not provide an endless scaling of problem size; FeMo-co is the largest enzyme cofactor in terms of the number of transition metals. Materials simulations provide a setting to scale the problem size, but one still faces the question as to whether the relevant states observed are truly that complicated classically. For example, classical simulations of the ground-state orders of the 2D Hubbard model currently show no exponential increase in difficulty when going to larger system sizes (Chen et al. 2025; Liu et al. 2025).
Has the availability of a classical strategy for FeMo-co changed your enthusiasm for quantum computers in chemistry?
Again, my answer is no. There is an entire community of nitrogenase scientists: experimental spectroscopists, synthetic chemists, and of course computational chemists, who are working to map out the reaction mechanism, none of whose research is predicated on using a quantum computer. Personally, I have never thought that to understand nitrogenase we would first have to build a quantum computer, otherwise I would not work on the problem!
At the same time, any computational tool brings new capabilities that
will be useful. Quantum algorithms come with theoretical guarantees; for
example, so long as the initial state is well prepared, we know the
error in the energy that we measure from a quantum algorithm, which is
more reliable than the classical estimates of error we obtain from
extrapolations. Similarly, initial state preparation for a quantum
computer, even for classically tractable problems, is probably easier
than solving the entire problem classically, since only a ‘rough’ guess
is needed. And finally, a polynomial or even constant factor speedup is
exciting, so long as the speedup is large enough!
Thus, I am in fact excited to see quantum computers applied to this
problem, I am just not waiting for them to be built first.
How should one think about past work on quantum algorithms that has used FeMo-co as a target?
FeMo-co was amongst the earliest examples of a chemical problem for which a case for quantum advantage was made. For this reason, it is overrepresented in the literature of quantum computing for chemistry. Should fully fault-tolerant quantum computers be available, they will naturally be applied to a wider set of systems (Chan 2024; Babbush et al. 2025).
Also, one must recognize that the availability of a single concrete optimization target has led to undeniable advances in quantum algorithms for quantum chemistry. In most cases, prior work to improve quantum resources estimates for FeMo-co involve techniques that apply to other systems as well. Thus, there’s no need to throw away those papers!
What are some lessons and conclusions to draw?
The first is obviously that, just because something has not been solved, or appears hard to solve classically, does not mean it is the best problem to choose for a quantum computer. The classical solution strategy for FeMo-co essentially involved a complicated classical state preparation problem, which is a shared challenge with ground-state estimation algorithms in quantum computers, and thus not perhaps an optimal choice of problem.
My second main conclusion is that since classical solutions in
complex problems are possible because they use some understanding of the
problem, for quantum algorithms to have maximum impact, they should use
the same knowledge. In fact most chemistry is not about truly mysterious
quantum systems, but more about ordinary quantum matter where we know
roughly what is going on, but where detailed simulations are still
required. If quantum computing algorithms can target this ‘mundane’
regime, they will have maximum impact on chemistry as it is practiced
today. In recent work, we have taken some steps in this direction by
proposing quantum algorithms for electronic structure that work within
the same heuristic framework as most current quantum chemistry
methods (Chen and
Chan 2025).
Finally, I wish to emphasize that, from the perspective of
understanding nitrogenase, and maximising societal impact, the choice of
computational algorithm and hardware to solve the problem is irrelevant.
The fact that FeMo-co electronic structure is not so mysterious is an
enormously positive thing, as it means that making progress on the
larger problem of the mechanism using computation no longer seems so
impossible. I have seen some of the brightest minds in the world helping
to advance quantum algorithms for this problem. If any of this
brainpower can be devoted to the chemical question itself, I believe we
can be very optimistic about the future solution of the nitrogenase
problem.
Arute, Frank, Kunal Arya, Ryan Babbush, et
al. 2019. “Quantum Supremacy Using a Programmable
Superconducting Processor.” Nature 574 (7779): 505–10.
Babbush, Ryan, Robbie King, Sergio Boixo, et al. 2025. “The Grand
Challenge of Quantum Applications.” arXiv Preprint
arXiv:2511.09124.
Badding, Edward D, Suppachai Srisantitham, Dmitriy A Lukoyanov, Brian M
Hoffman, and Daniel LM Suess. 2023. “Connecting the Geometric and
Electronic Structures of the Nitrogenase Iron–Molybdenum Cofactor
Through Site-Selective 57Fe Labelling.”
Nature Chemistry 15 (5): 658–65.
Berry, Dominic W, Craig Gidney, Mario Motta, Jarrod R McClean, and Ryan
Babbush. 2019. “Qubitization of Arbitrary Basis Quantum Chemistry
Leveraging Sparsity and Low Rank Factorization.” Quantum
3: 208.
Chan, Garnet Kin-Lic. 2024. “Spiers Memorial Lecture: Quantum
Chemistry, Classical Heuristics, and Quantum Advantage.”
Faraday Discussions 254: 11–52.
Chen, Ao, Zhou-Quan Wan, Anirvan Sengupta, Antoine Georges, and
Christopher Roth. 2025. “Neural Network-Augmented Pfaffian
Wave-Functions for Scalable Simulations of Interacting Fermions.”
arXiv Preprint arXiv:2507.10705.
Chen, Jielun, and Garnet Kin Chan. 2025. “A Framework for Robust
Quantum Speedups in Practical Correlated Electronic Structure and
Dynamics.” arXiv Preprint arXiv:2508.15765.
Einsle, Oliver, and Douglas C Rees. 2020. “Structural Enzymology
of Nitrogenase Enzymes.” Chemical Reviews 120 (12):
4969–5004.
Gray, Johnnie, and Stefanos Kourtis. 2021. “Hyper-Optimized Tensor
Network Contraction.” Quantum 5: 410.
Jiang, Hao, and Ulf Ryde. 2023. “N2 Binding to the E0–E4 States
of Nitrogenase.” Dalton Transactions 52 (26): 9104–20.
Lancaster, Kyle M, Michael Roemelt, Patrick Ettenhuber, et al. 2011.
“X-Ray Emission Spectroscopy Evidences a Central Carbon in the
Nitrogenase Iron-Molybdenum Cofactor.” Science 334
(6058): 974–77.
Lee, Seunghoon, Joonho Lee, Huanchen Zhai, et al. 2023.
“Evaluating the Evidence for Exponential Quantum Advantage in
Ground-State Quantum Chemistry.” Nature Communications
14 (1): 1952.
Li, Zhendong, Sheng Guo, Qiming Sun, and Garnet Kin-Lic Chan. 2019.
“Electronic Landscape of the P-Cluster of Nitrogenase as Revealed
Through Many-Electron Quantum Wavefunction Simulations.”
Nature Chemistry 11 (11): 1026–33.
Liu, Wen-Yuan, Huanchen Zhai, Ruojing Peng, Zheng-Cheng Gu, and Garnet
Kin-Lic Chan. 2025. “Accurate Simulation of the Hubbard Model with
Finite Fermionic Projected Entangled Pair States.” Physical
Review Letters 134 (25): 256502.
Lovell, Timothy, Jian Li, Tiqing Liu, David A Case, and Louis Noodleman.
2001. FeMo Cofactor of
Nitrogenase: A Density Functional Study of States MN, MOX, MR, and
MI.” Journal of the American Chemical Society 123 (49):
12392–410.
Low, Guang Hao, Robbie King, Dominic W Berry, et al. 2025. “Fast
Quantum Simulation of Electronic Structure by Spectrum
Amplification.” arXiv Preprint arXiv:2502.15882.
Luo, Maxine, and J Ignacio Cirac. 2025. “Efficient Simulation of
Quantum Chemistry Problems in an Enlarged Basis Set.” PRX
Quantum 6 (1): 010355.
Reiher, Markus, Nathan Wiebe, Krysta M Svore, Dave Wecker, and Matthias
Troyer. 2017. “Elucidating Reaction Mechanisms on Quantum
Computers.” Proceedings of the National Academy of
Sciences 114 (29): 7555–60.
Sharma, Sandeep, Kantharuban Sivalingam, Frank Neese, and Garnet Kin-Lic
Chan. 2014. “Low-Energy Spectrum of Iron–Sulfur Clusters Directly
from Many-Particle Quantum Mechanics.” Nature Chemistry
6 (10): 927–33.
Thorhallsson, Albert Th, Bardi Benediktsson, and Ragnar Bjornsson. 2019.
“A Model for Dinitrogen Binding in the E4 State of
Nitrogenase.” Chemical Science 10 (48): 11110–24.
Thorneley, Roger NF, and DJ Lowe. 1984. “The Mechanism of
Klebsiella Pneumoniae Nitrogenase Action. Pre-Steady-State Kinetics of
an Enzyme-Bound Intermediate in N2 Reduction and of NH3
Formation.” Biochemical Journal 224 (3): 887–94.
Wan, Kianna, Mario Berta, and Earl T Campbell. 2022. “Randomized
Quantum Algorithm for Statistical Phase Estimation.” Physical
Review Letters 129 (3): 030503.
Yamaguchi, Kizashi, Takayuki Fueno, Masa-aki Ozaki, Norikazu Ueyama, and
Akira Nakamura. 1990. “A General Spin-Orbital (GSO) Description of
Antiferromagnetic Spin Couplings Between Four Irons in Iron-Sulfur
Clusters.” Chemical Physics Letters 168 (1): 56–62.
Zhai, Huanchen, Seunghoon Lee, Zhi-Hao Cui, Lili Cao, Ulf Ryde, and
Garnet Kin-Lic Chan. 2023. “Multireference Protonation Energetics
of a Dimeric Model of Nitrogenase Iron–Sulfur Clusters.” The
Journal of Physical Chemistry A 127 (47): 9974–84.
Zhai, Huanchen, Chenghan Li, Xing Zhang, Zhendong Li, Seunghoon Lee, and
Garnet Kin-Lic Chan. 2026. “Classical Solution of the FeMo-Cofactor
Model to Chemical Accuracy and Its Implications.” arXiv
Preprint arXiv:2601.04621.